Defining
1st-level generative AI as applications that are directly based on X-to-Y models (foundation models that build a kind of operating system for downstream tasks) where X and Y can be text/code, image, segmented image, thermal image, speech/sound/music/song, avatar, depth, 3D, video, 4D (3D video, NeRF), IMU (Inertial Measurement Unit), amino acid sequences (AAS), 3D-protein structure, sentiment, emotions, gestures, etc., e.g.
- X = text, Y = text: LLM-based chatbots like ChatGPT (from OpenAI based on LLMs GPT-3.5 [4K context] or GPT-4 [8K/32K context]), Bing Chat (GPT-4), Bard (from Google, based on PaLM 2), Claude (from Anthropic [100K context]), Llama2 (from Meta), Falcon 180B (from Technology Innovation Institute), Alpaca, Vicuna, OpenAssistant, HuggingChat (all based on LLaMA [GitHub] from Meta), OpenChatKit (based on EleutherAI’s GPT-NeoX-20B), CarperAI, Guanaco, My AI (from Snapchat), Tingwu (from Alibaba based on Tongyi Qianwen), (other LLMs: MPT-7B and MPT-30B from Mosaic [65K context, commercially usable], Orca, Open-LLama-13b), or coding assistants (like GitHub Copilot / OpenAI Codex, AlphaCode from DeepMind, CodeWhisperer from Amazon, Ghostwriter from Replit, CodiumAI, Tabnine, Cursor, Cody (from Sourcegraph), StarCoder from Big Code Project led by Hugging Face, CodeT5+ from Salesforce, Gorilla, StableCode from Stability.AI, Code Llama from Meta), or writing assistants (like Jasper, Copy.AI), etc.
- X = text, Y = image: Dall-E (from OpenAI), Midjourney, Stable Diffusion (from Stability.AI), Adobe Firefly, DeepFloyd-IF (from Deep Floyd, [GitHub, HuggingFace]), Imagen and Parti (from Google), Perfusion (from NVIDIA)
- X = text, Y = 360° image: Skybox AI (from Blockade Labs)
- X = text, Y = 3D avatar: Tafi
- X = text, Y = avatar lip sync: Ex-Human, D-ID, Synthesia, Colossyan, Hour Once, Movio, YEPIC-AI, Elai.io
- X = speech + face video, Y = synched audio-visual: Lalamu
- X = text, Y = video: Gen-2 (from Runway Research), Imagen-Video (from Google), Make-A-Video (from Meta), or from NVIDIA
- X = text, Y = video game: Muse & Sentis (from Unity)
- X = image, Y = text: GPT-4 (from OpenAI), LLaVA
- X = image, Y = segmented image: Segment Anything Model (SAM by Meta)
- X = speech, Y = text: STT (speech-to-text engines) like Whisper (from OpenAI), MMS [GitHub] (from Meta), Conformer-2 (from AssemblyAI)
- X = text, Y = speech: TTS (text-to-speech engines) like VALL-E (from Microsoft), Voicebox (from Meta), SoundStorm (from Google), ElevenLabs, Bark, Coqui
- X = text, Y = music: MusicLM (from Google), RIFFUSION, AudioCraft (MusicGen, AudioGen, EnCodec from Meta), Stable Audio (from Stability.ai)
- X = text, Y = song: Voicemod
- X = text, Y = 3D: DreamFusion (from Google)
- X = text, Y = 4D : MAV3D (from Meta)
- X = image, Y = 3D : CSM
- X = image, Y = audio: ImageBind [1] (from Meta, on GitHub)
- X = audio, Y = image: ImageBind [2] (from Meta)
- X = music, Y = image: MusicToImage
- X = text, Y = image & audio: ImageBind [3] (from Meta)
- X = audio & image, Y = image: ImageBind [4] (from Meta)
- X = IMU, Y = video: ImageBind (from Meta)
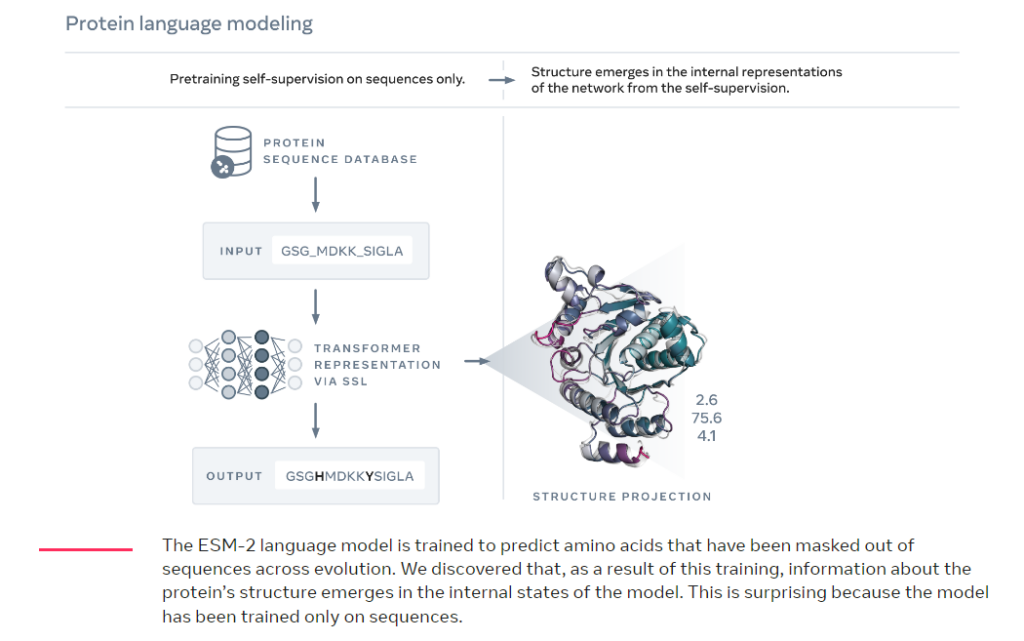
- X = AAS, Y = 3D-protein: AlphaFold (from Google), RoseTTAFold (from Baker Lab), ESMFold (from Meta)
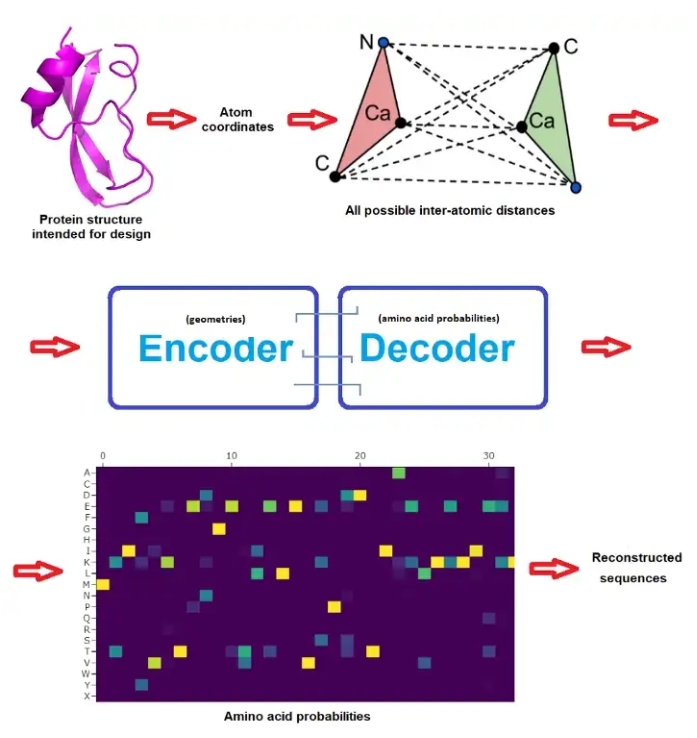
- X = 3D-protein, Y = AAS: ProteinMPNN (from Baker Lab)
- X = 3D structure, Y = AAS: RFdiffusion (from Baker Lab)
and 2nd-level generative AI that builds some kind of middleware and allows to implement agents by simplifying the combination of LLM-based 1st-level generative AI with other tools via actions (like web search, semantic search [based on embeddings and vector databases like Pinecone, Chroma, Milvus, Faiss], source code generation [REPL], calls to math tools like Wolfram Alpha, etc.), by using special prompting techniques (like templates, Chain-of-Thought [COT], Self-Consistency, Self-Ask, Tree Of Thoughts, ReAct [Reason + Act], Graph of Thoughts) within action chains, e.g.
- ChatGPT Plugins (for simple chains)
- LangChain + LlamaIndex (for simple or complex chains)
- ToolFormer
we currently (April/May/June 2023) see a 3rd-level of generative AI that implements agents that can solve complex tasks by the interaction of different LLMs in complex chains, e.g.
- BabyAGI
- Auto-GPT
- Llama Lab (llama_agi, auto_llama)
- Camel, Camel-AutoGPT
- JARVIS (from Microsoft)
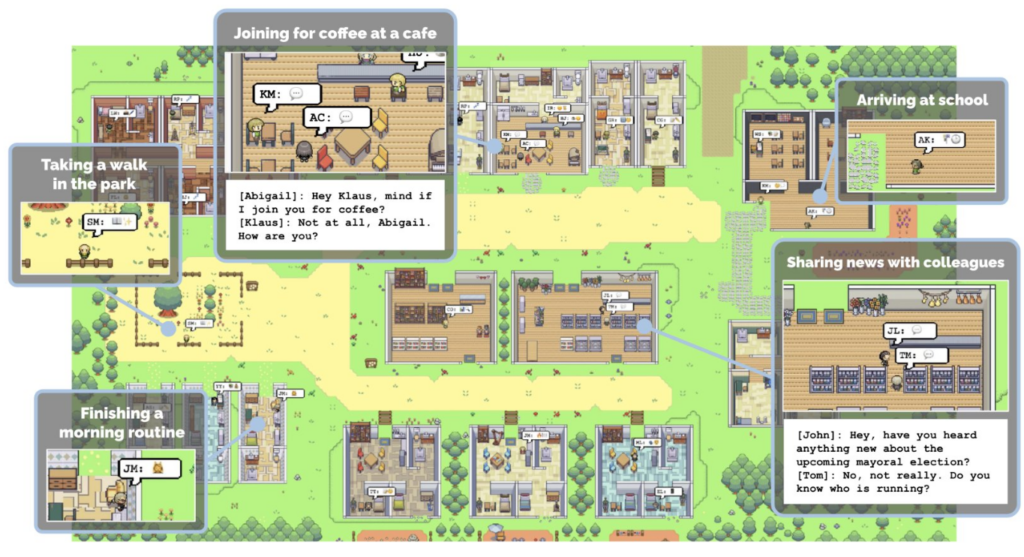
- Generative Agents
- ACT-1 (from Adept)
- Voyager
- SuperAGI
- GPT Engineer
- Parsel
- MetaGPT
However, older publications like Cicero may also fall into this category of complex applications. Typically, these agent implementations are (currently) not built on top of the 2nd-level generative AI frameworks. But this is going to change.
Other, simpler applications that just allow semantic search over private documents with a locally hosted LLM and embedding generation, such as e.g. PrivateGPT which is based on LangChain and Llama (functionality similar to OpenAI’s ChatGPT-Retrieval plugin), may also be of interest in this context. And also applications that concentrate on the code generation ability of LLMs like GPT-Code-UI and OpenInterpreter, both open-source implementations of OpenAI’s ChatGPT Code Interpreter/AdvancedDataAnalysis (similar to Bard’s implicit code execution; an alternative to Code Interpreter is plugin Noteable), or smol-ai developer (that generates the complete source code from a markup description) should be noticed.
There is a nice overview of LLM Powered Autonomous Agents on GitHub.
The next level may then be governed by embodied LLMs and agents (like PaLM-E with E for Embodied).